Genome methylation patterns in male breast cancer – Identification of an epitype with hypermethylation of polycomb target genes
Johansson I, Lauss M, Holm K, Staaf J, Nilsson C, Fjällskog ML, Ringnér M, Hedenfalk I
Mol Oncol. 2015 Oct;9(8):1565-79. doi: 10.1016/j.molonc.2015.04.013. Epub 2015 May 8
Background
The question of whether male breast cancers are the same as female breast cancers can be, and must be asked at many levels of analysis. Biology is complex.
Take for example the way that DNA, the substance that encodes our genes, works. At its simplest, DNA encodes RNA which encodes protein: a single copy of a gene in the DNA can make many copies in RNA which can in turn be used by the cell to make many copies of the protein it encodes.
One level of information in the genome is therefore its sequence: the letters that are written there, and their order. Another level of information, also inherited from cell to cell, and in a different way from parent to offspring, is so-called epigenetic, meaning a layer above the genetic sequence. One type of epigenetic information is encoded by DNA methylation – the decoration of DNA by chemical modifcations consisting of methyl groups. Think “track changes” annotations in a word document. There is a whole ecosystem of enzymes whose job it is to put methyl groups onto DNA, read what methyl groups are there and remove them. In general the more methylated a region of DNA is, the less active the genes encoded by it are. One of the main roles of DNA methylation is to program the genome for different tissues by putting away genes not needed for a given cell type, and making those that are needed more active.
In cancer, DNA methylation can sometimes go wrong, and it can lead to the inappropriate inactivation of genes, helping the cancer grow. In many cancers genome-wide analysis of DNA methylation has led to the identification of sub-types that share a particular methylation pattern, related to a particular biology. The question that Johansson and colleagues set out to answer was whether male breast cancer had such sub-groups, and if so, whether they resembled those previously identified for female breast cancer.
Findings
The take home message from Johansson’s analysis of DNA methylation of 47 male breast cancers is that while overall they do resemble the same histological type of female cancers, there are some differences. This is beginning to sound familiar to my regular readers, I am sure – we have seen it at other levels of analysis, including that of mutations in the cancer genome just recently.
Let’s look in some more detail. Johansson and colleagues analyzed genome wide methylation using arrays, a technology that allows the efficient survey of the entire genome. They identified two so called ‘epitypes’, ME1 and ME2 into which the male tumors clustered, based on the patterns observed. Here is a summary of their findings for these two groups:
| ME1 | ME2 | |
|---|---|---|
| previously identified transcriptional subgroup they are associated with | luminal M1 | luminal M2 |
| estrogen receptor activity associated gene expression | less | more |
| tumor growth rate | higher | lower |
| genome instability | higher | lower |
| methylation pattern | quite different from normal cells; HOX genes targeted; decreased expression of PRC2 target genes; higher levels of EZH2, a master regulator of methylation | more similar to normal cells |
| summary | more aggressive looking tumors | less aggressive looking tumors |
The identification of subgroups based on a molecular signature is now a quite common observation for most cancers: tumors that may look the same down the microscope and so may receive identical histopathological diagnoses, differ at the molecular level, revealing additional nuances of their biology. Ultimately these may allow precision medicine approaches that provide more effective treatments for the more aggressive cancers that are less likely to benefit from conventional therapy. The fact that the epigenetic analysis in this paper agrees with the gene expression analysis done previously by this team is expected – after all the two are linked directly by how DNA methylation influences how the genome works (see above).
So what about comparing the men and the women? Well, to the credit of this group of investigators the same methodology was used for the men here, and a larger group of women previously, and so a direct comparison was possible. Using a mathematical strategy that puts tumors with a similar methylation pattern across thousands of regions together in a so called cluster, the authors identified three main groups:
- basal female breast cancers
- luminal B female breast cancers and ME1 male breast cancers
- luminal A female breast cancers and ME2 male breast cancers
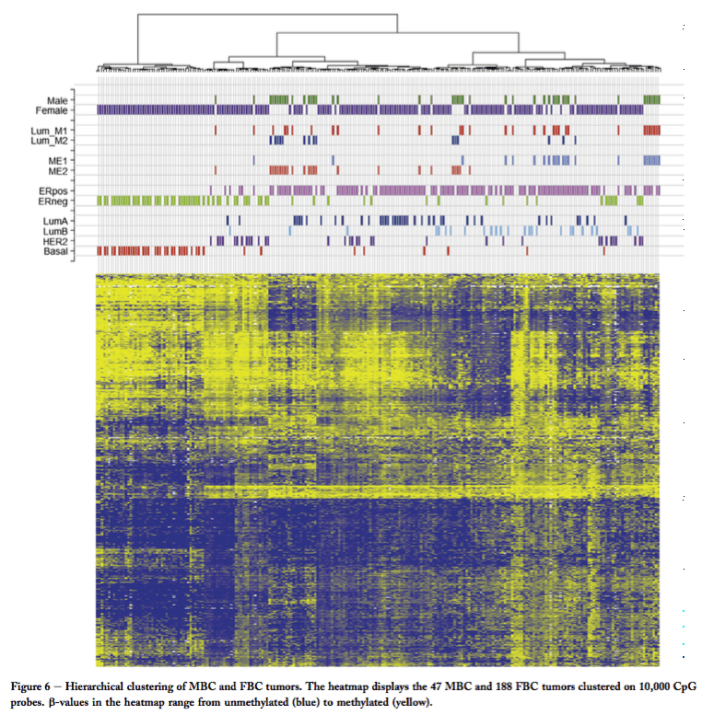
Hierarchical clustering of male and female breast cancers – Fig. 6 from Johansson et al (http://www.ncbi.nlm.nih.gov/pubmed/25990542)
Luminal A and B are groups previously identified by genome analysis of women’s breast cancers. So, the finding is that overall male breast cancers are similar to female breast cancers. Furthermore, as expected almost all male breast cancers associate with the luminal groups which are ER positive, and very few with the basal which is hormone-independent. So, overall, breast cancers in men and women show similar groupings.
But, when you look in more detail, within luminal clusters, the male cases tend to stick together. Imagine a large family that share an overall physical resemblance. Within it, you’d still see smaller groups that look even more alike, say siblings rather than cousins. This suggests that the male breast cancers may have a few shared characteristics not shared by the overall group they belong to – these are listed in the table. Interestingly a similar phenomenon was also the case for some of the aspects of exactly where the methylation was observed relatively to gene stucture.
Comment
A striking pattern is emerging in the molecular analysis of male breast cancers. They are clearly breast cancers, sharing the majority of their characteristics with the same class of female tumors, no matter how you analyze them: histologically, by gene mutation and now epigenetically. However, the male breast cancers also have some aspects of their detailed biology that are somewhat distinct.
I would argue that in the fading age of “chemo-surgery-radiation-endocrine” therapy these subtle molecular differences may not have been all that important. However, in the dawning era of precision medicine they may well be the entire ball game.

Absolutely, very interesting and important findings…I’m sure there is more to come. Thanks for sharing this information.